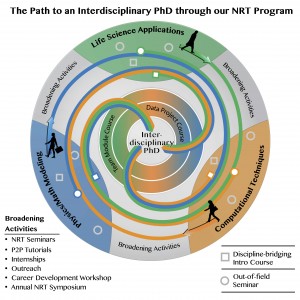
COMBINE fellows are generally most involved in program components during a 2 year period within their graduate studies. They will typically engage in introductory disciple-bridging course work in their first year and undertake the most intensive program components in their second year (see sample timeline). Fellows will continue involvement throughout the remainder of their graduate careers, with fellows in their later years of study serving as mentors to those in their early years.
COMBINE students are drawn from three different disciplines:
- Life Sciences: from UMD graduate programs in: Behavior, Ecology, Evolution, and Systematics; Molecular and Cellular Biology; Computational Biology, Bioinformatics and Genomics; Neuroscience and Cognitive Sciences; Epidemiology; and Biostatistics and Bioinformatics
- Physical and Mathematical Sciences: from UMD graduate programs in Physics, Biophysics, and Applied Mathematics and Scientific Computing
- Computational Sciences: from UMD graduate programs in Computer Science as well as Electrical and Computer Engineering
Required Core COMBINE Components: The following represent the program components required for COMBINE fellows. Students who complete the the 12 credits of required coursework will earn a COMBINE Graduate Certificate alongside their PhD. See sample timeline for a typical progression of COMBINE components.
- Discipline-bridging coursework (4 credits): In addition to completing their PhD degree requirements in one of three disciplinary areas, COMBINE fellows must complete: (A) one regular (3+ credit) course at the graduate or advanced undergraduate level from one of the other two disciplines (chosen from a list of appropriate courses) and (B) one out-of-field graduate seminar course (1+ credit) from the third discipline. This introductory coursework is designed to help bridge the physical/mathematical, computational, and life sciences. For example, physics/math and computer science graduate students gain familiarity with core concepts in the life sciences (via courses such as BIOL708: Cell Biology from a Biophysical Perspective or BSCI453: Cellular Neurophysiology), life science and computer science graduate students study the techniques of physical and mathematical modeling (via courses such as PHYS615: Nonlinear Dynamics of Extended Systems or MATH420: Mathematical Modeling), and life and physical science graduate students explore computational techniques for processing and analyzing large datasets (via courses such CBMG688: Programming for Biologists or AMSC660: Scientific Computing). For students in programs outside the computational sciences who have no prior computational training, we require that the 3+ credit course be a computation-based course. Note that some courses fulfilling these “out-of-discipline” requirements may actually be taught from within the student’s primary discipline. For example, the course BSCI474: Mathematical Biology may be used by a life science graduate student to fulfill the COMBINE course requirement for physical/mathematical modeling. List of Approved Discipline-Bridging Courses
- Advanced Interdisciplinary coursework: Computational and Mathematical Analysis of Biological Networks across Scales (CMSC828O)(3 credits; Syllabus): Typically in their second year of graduate study, COMBINE students undertake advanced interdisciplinary coursework in the form of a team-based module course. This innovative course was developed and has been taught by Prof. Hector Corrada Bravo. The central idea of this course is that students, working in small interdisciplinary teams, learn network science methodologies and apply them to large, complex datasets drawn from life science applications. Modules are organized around the four major methodological themes of our NRT program: network measures, mechanistic network models, network statistics and machine learning, and network visualization.
- Advanced Interdisciplinary coursework: Interdisciplinary Research and Communication Practicum for Data-Driven Science(PHYS782, formerly PHYS798N)) (3 credits; Syllabus): COMBINE students participate in a semester-long intensive data project, to be guided by one or more of our COMBINE faculty and/or partners. The motivating idea behind this course is to fill a major gap in graduate science education by helping students develop and hone the skills necessary for conducting ground-breaking research, instead of focusing solely on building a knowledge base. In addition, the course has a significant focus on developing skills for communication to diverse audiences. Students learn to write for and speak to individuals in the same field, individuals in another specified field to which their research is applicable, and a general science audience.
- COMBINE seminars (2 semesters required, 1 credit each): Each semester, we hold a weekly COMBINE seminar series. In the Fall, the seminar (PHYS780, formerly PHYS798T) takes on a reading group format and trainees discuss seminal papers in network biology (see Fall COMBINE Seminar Syllabus). In the Spring, the seminar (PHYS781, formerly PHYS798U) focuses on research-in-progress (see Spring COMBINE Seminar Syllabus). In both semesters, students have opportunities to develop their oral communication skills. Both seminars also feature a few invited presentations from faculty who describe their research and offer career advice.
- Peer-to-peer (P2P) tutorials (week-long event held between semesters): This event is held twice a year: in January between the Fall and Spring semesters and in the summer before the start of the Fall semester. Students work in small groups of 3-4 preparing tutorials for their peers. Groups and topics are chosen by the trainees themselves, with each group advised by one or more COMBINE faculty members of its choosing. Each fellow must participate in the development of one P2P tutorial during the course of their studies. Topics have included: intro to machine learning, tools for reproducible science, and mentoring undergraduate research.
- Out of field mentor: We require every COMBINE fellow to identify an out-of-field mentor by the end of their second semester of participation in the program, or earlier if possible. At the start of the program, students are categorized in one of 3 broad domains: 1) life sciences, 2) physical and mathematical sciences 3) computational science and engineering. The out-of-field mentor should be a faculty member (or senior researcher at another institution) from outside the fellow’s domain. For example, a student from the life sciences needs to identify an out-of-field mentor from category 2 or 3. Ideally, the out-of-field mentor would be a faculty member who is part of the COMBINE program or is interested in joining the COMBINE faculty. The role of the out-of-field mentor is to consult with the student regarding his or her path through the COMBINE program and offer advice about coursework and research. At a minimum, COMBINE fellows must meet with their out-of-field mentors by the end of their first year in the program and again before the end of their second year. The fellow and the out-of-field mentor will jointly fill out a short progress form highlighting strengths and outlining strategies for improvement. We encourage students to pursue a research collaboration with their out-of-field mentors where feasible and appropriate. The out-of-field mentor is expected to serve as a member of the student’s thesis committee. You should discuss with your Adviser who this out-of-field mentor could be, with the goal being to incentivize cross-disciplinary collaboration.Fellows whose research plan changes during the course of their study can change their out-of-field mentor when appropriate, with approval from the COMBINE Director.
Additional Broadening Activities:
- Internship opportunities with COMBINE partners (optional): COMBINE leverages strong existing partnerships between UMD and four nearby centers for research excellence: the Smithsonian Institution, where students can explore evolutionary, ecological, and conservation problems; the National Institutes of Health, where students can study the roles of biomolecular and neuronal networks in disease; the National Institute of Standards and Technology, where students can use network science to develop new approaches for bioscience measurement and interpretation; and the University of Maryland School of Medicine, where students can apply network analysis techniques in the context of health informatics and bioimaging. In addition to providing internship opportunities with these local research partners, our program offers students the opportunity to explore alternate career paths via internships with industry and government partners who are invested in the translation of scientific advancements to a range of applications.
- Outreach and peer mentoring (required): COMBINE students actively participate in outreach and mentoring activities to promote cross-disciplinary STEM education to undergraduate and middle/high school groups. A principal goal is to promote participation from groups traditionally under-represented in STEM fields. Outreach and mentoring activities include: outreach to middle/high school students and the general public via UMD channels like CompSci Connect and GEMS; undergraduate research mentoring; peer-mentoring in which fellows meet 2-3 times per year in 3-person peer mentoring teams with representation from different disciplines and graduate career stages. Fellows are required to engage in peer-mentoring and at least one outreach activity during their participation in COMBINE.
- Annual Career Develop Workshop: Each year, we hold a Career Development Workshop for our trainees that brings in a diverse set of speakers to discuss how network-focused data science methods can be applied in a variety of career contexts. The workshop is largely student-organized with NRT faculty members providing overall guidance. The workshop’s theme changes from year to year, depending on student interests. Speakers are drawn from our established industry and government partners, as well from other organizations who might wish to become involved in our program.
- Annual COMBINE Symposium: Every year, we host a one-day Annual COMBINE Symposium to showcase our research activities. The symposium format includes oral and poster presentations by COMBINE fellows, presentations from leading researchers outside of our NRT program, as short talks from some of the COMBINE faculty, and panels.